 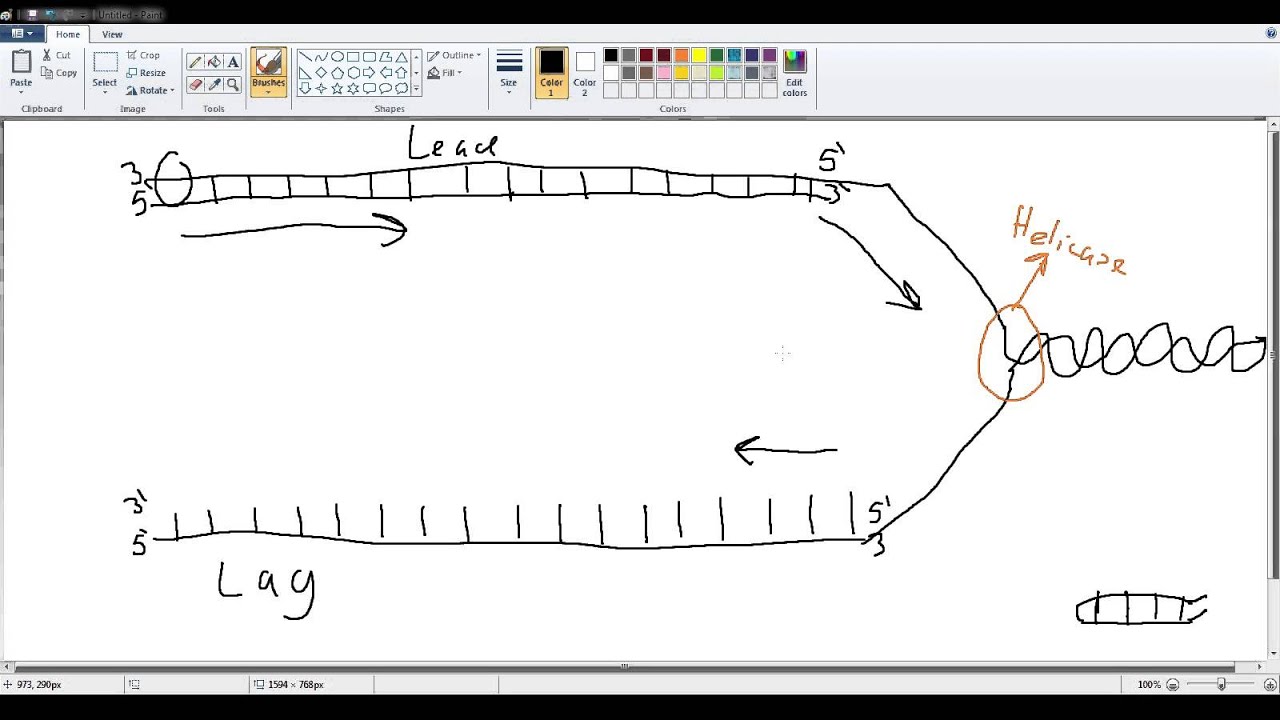
L'analyse comparative des profils OFR montre que les changements réplicatifs permettent la classification des lignées cellulaires en fonction de leur tissu d'origine, la nature cancéreuse ou non de la lignée n'intervenant qu'en second ordre. Nous proposons une caractérisation originale de la plasticité du programme de réplication de l’ADN se basant sur le profilage de 12 lignées cellulaires humaines normales ou cancéreuses par la méthode Ok-seq de purification et séquençage des fragments d'Okazaki qui permet de déterminer l'orientation de la progression des fourches de réplication (OFR) à haute résolution (10 kilo bases). 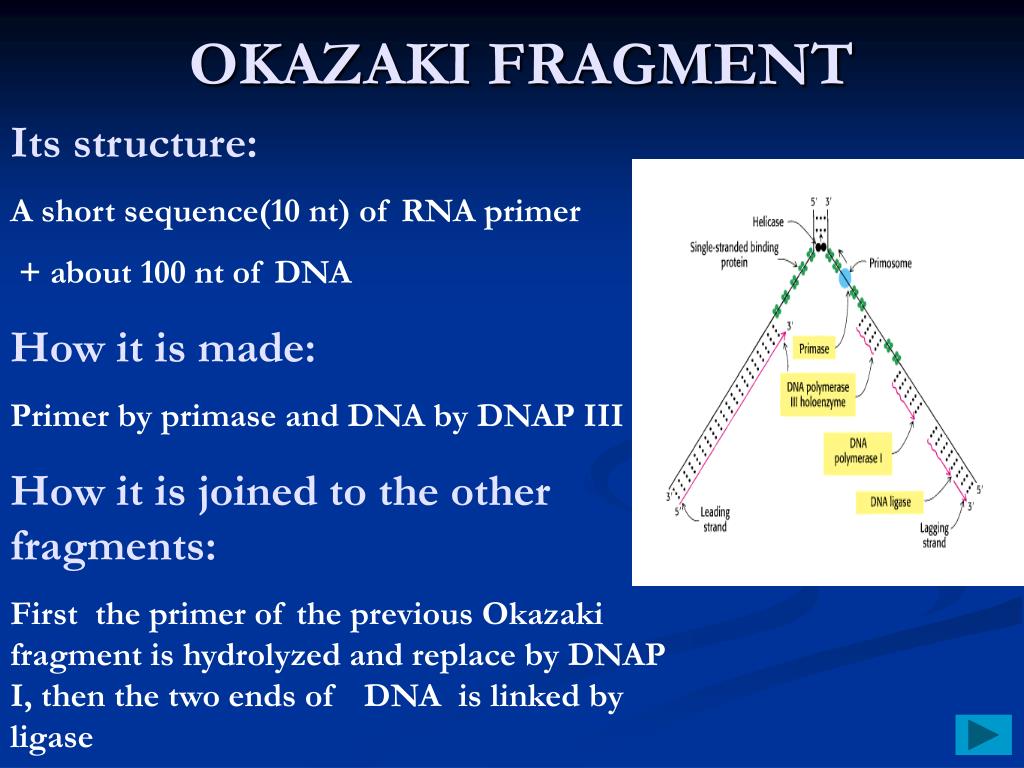
Le programme spatiotemporel de réplication de l'ADN est régulé au cours du développement et altéré durant la progression cancéreuse. This work provides an original overview of replication program changes during normal or pathological differentiation, including a cell line specific control of IZ in late-replication gene deserts. This leads us to quantify the degree of dissociation between IZ and activation of transcription. In contrast, the late replicated, low gene density and low G+C content regions have few efficient IZs, often specific to a tissue or lineage. Nevertheless, the G+C rich and active gene regions, replicated early in the S phase, have the most stable replication program, they present a high density of efficient replication initiation zones (IZ) conserved between cell lines. 
There is no hotspot for the accumulation of replicative changes, they are widely dispersed throughout the genome. Comparative analysis of the RFD profiles shows that the replicative changes allow the classification of the cell lines according to their tissue of origin, the cancerous or non-cancerous nature of the cell line type intervening only in second order. We propose an original characterization of the plasticity of the DNA replication program based on the profiling of 12 normal or cancerous human cell lines by the Ok-seq method of purification and sequencing of Okazaki fragments which allows to determine the orientation of the progression of replication forks (RFD) at high resolution (10 kilo bases). Sites of termination are, therefore, indicative of origin firing time – allowing us to faithfully reconstruct the temporal dynamics of the replication program using data from a single asynchronous culture.The spatiotemporal program of DNA replication is regulated during development and altered during cancer progression. Contrary to expectation, we find that centromeres and highly transcribed regions are not strong determinants of replication termination rather, termination generally occurs midway between two adjacent replication origins at positions dictated by the relative firing times of those origins. We provide evidence that S-phase follows a distinct temporal program dominated by replication origins firing with high probability within distinct time intervals. Our methodology also provides a genome-wide view of DNA replication: allowing the first detailed measurements of the efficiencies of all replication origins and regions of termination. Using deep sequencing, we have found that Okazaki fragment synthesis is strongly influenced by nucleosomes as well as certain other DNA bound factors. These Okazaki fragments are typically less than 500bp in length and are generally sized according to the nucleosome repeat. Recently, using transient DNA ligase I inactivation, we have been able to capture and analyze short DNA molecules produced on the lagging strand. Synthesis of the lagging strand occurs discontinuously and necessitates the repeated production of Okazaki fragments. DNA replication is inherently asymmetric. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
